Introduction
Wikipedia has a pretty good definition of workflow: an orchestrated and repeatable pattern of activity, enabled by the systematic organization of resources into processes that transform materials, provide services, or process information. Doing Science involves the workflow of repeating and documenting experiments and the related data analysis. Consequently, managing workflow is critically important for the scientific endeavor. There have been dozens of projects that have built tools to simplify the process of specifying and managing workflow in science. We described some of these in our 2007 book “Workflows for e-Science” and another Wikipedia article gives another list of over a dozen scientific workflow system and this page lists over 200 systems. Many of these systems were so narrowly focused on a single scientific domain or set of applications that they have not seen broad adoption. However, there are a few standouts and some of these have demonstrated they can manage serious scientific workflow.
Pegasus from ISI is the most well-known and used framework for managing science workflow. To use it on the cloud you deploy a Condor cluster on a set of virtual machines. Once that is in place, Pegasus provides the tools you need to manage large workflow computations. Makeflow from Notre Dame University is another example. Based on a generalization of the execution model for the old Unix Make system, Makeflow uses Condor but it also has a native distributed platform called WorkQueue. Makeflow is commonly used on large HPC Supercomputers but they also claim an implementation on AWS Lambda.
Workflow in the Cloud.
Doing scientific computing in the cloud is different from the traditional scientific data centers built around access to a large supercomputer. A primary difference is that the cloud support models of computing not associated with traditional batch computing frameworks. The cloud is designed to support continuously running services. These may be simple web services or complex systems composed of hundreds of microservices. The cloud is designed to scale to your needs (and budget) on-demand. The largest commercial public clouds from Amazon, Google and Microsoft are based on far more than providing compute cycles. They offer services to build streaming applications, computer vision and speech AI services, scalable data analytics and database services, managing edge devices, robotics and now attached quantum processors. While there are tools to support batch computing (Microsoft Azure even has an attached Cray), the cloud is also an excellent host for interactive computational experimentation.
Christina Hoffa, et. al. “On the Use of Cloud Computing for Scientific Workflows” describe some early 2008 experiments using cloud technology for scientific workflow. The cloud of 2019 presents many possibilities they did not have access to. Two of these are “cloud native” microservice frameworks such as Kubernetes and serverless computing models.
Kubernetes has been used for several workflow systems. An interesting example is Reana from CERN. Reana is a research data analysis platform that runs on your desktop or on a cloud Kubernetes cluster. Reana uses several workflow languages but the one that is most frequently used is CWL, the Common Workflow Language, which is rapidly becoming an industry standard. CWL is used in a number of other cloud workflow tools including AVADOS from Veritas Genetics, a version of the popular Apache Airflow workflow tools and several other systems with implementations “in progress”. Argo is another workflow took that is designed to work with Kubernetes.
Workflow Using Serverless Computing
Serverless computing is a natural fit for workflow management. Serverless allows applications to run on demand without regard to compute resource reservation or management. Serverless computations are triggered by events. Typical among the list of event types are:
- Messages arriving on Message Queues
- Changes in Databases
- Changes in Document Stores
- Service APIs being invoked
- Device sensor data sending new updates
These event types are each central to workflow automation. AWS Lambda was the first production implementation of a serverless platform, but not the last. Implementations from Microsoft, IBM and Google are now available and the open source implementation from OpenWhisk is available for OpenStack and other platforms.
Serverless computing is built around the concept of “function as a service” where the basic unit of deployment is not a VM or container, but the code of a function. When the function is deployed it is tied to a specific event category such as one of those listed above. These functions are intended to be relatively light weight (not a massive memory footprint and a short execution time). The semantics of the function execution dictate that they are stateless. This allows many instances of the same function to be responding to events at the same time without conflict. The function instances respond to an event, execute and terminate. While the function itself is stateless, it can affect the state of other object during its brief execution. It can write files, modify databases, and invoke other functions.
Workflow State Management
Most scientific workflows can be described as a directed acyclic graph where the nodes are the computational steps. An arc in the graph represents a completion of a task that signals another it may start. For example, the first task writes a file in a storage container and that triggers an event which fires the subsequent task that is waiting for the data in the file. If the graph takes the shape of a tree where one node creates events which trigger one or more other nodes, the translation to serverless is straightforward: each node of the graph can be compiled into one function. (We show an example of this case in the next section.)
One of the challenges of using serverless computing for workflow is state management. If the “in degree” of a node is greater than one, then it requires more than one event to trigger the event. Suppose there are two events that must happen before a node is triggered. If the function is stateless it cannot remember that the one of the conditions has already been met. The problem is that the graph itself has state defined by which nodes have been enabled for execution. For example, Figure 1 is a CWL-like specification of such a case. NodeC cannot run until NodeA and NodeB both complete.

Figure 1. A CWL-like specification of a three step workflow where the third step requires that both the first step and second step are complete. The first and second step can run simultaneously or in any order.
One common solution to this problem is to assume there is a persistent, stateful instance of a workflow manager that holds the graph and keeps track of its global state. However, it is possible to manage the workflow with a single stateless function. To see how this can be done notice that in the above specification each of the step nodes requires the existence of one or more input files and the computation at that node produces one or more output files. As shown in Figure 2 below, workflow stateless function listens to the changes to the file system (or a database).

Figure 2. A workflow manager/listener function responds to events in the file system created by the execution of the applications. As shown in the next session, if the app is small, it may be embedded in the manager, but otherwise it can be containerized and run elsewhere in the cloud.
Here we assume that the application invocations, which are shown as command-line calls in Figure 1, are either executed in the lambda function or by invocations to the application wrapped as a Docker container running as a microservice in Kubernetes. When an application terminates it deposits the output file to the file system which triggers an event for the workflow manager/listener function.
The workflow manager/listener must then decide which step nodes were affected by this event and then verify that all the conditions for that node are satisfied. For example, if the node requires two file, it much check that both are there before invoking the associated application. There is no persistent state in the manager/listener as it is all part of the file system. To run multiple workflow instances concurrently each event and file must have an instance number ID as part of its metadata.
A Simple Example of Workflow using AWS Lambda
In the following paragraphs we describe a very simple implementation of a document classification workflow using AWS Lambda. The workflow is a simple tree with two levels and our goal here is to demonstrate the levels of concurrency possible with a serverless approach. More specifically we demonstrate a system that looks for scientific documents in an AWS S3 bucket and classifies them by research topic. The results are stored in an Amazon DynamoDB table. The documents each consist of a list of “scientific phrases”. An example document is below.
“homology is part of algebraic topology”,
‘The theory of evolution animal species genes mutations’,
“the gannon-lee singularity tells us something about black holes”,
‘supercomputers are used to do very large simulations’,
‘clifford algebras and semigroup are not fields and rings’,
‘galaxies have millions of stars and some are quasars’,
‘deep learning is used to classify documents and images’,
‘surgery on compact manifolds of dimension 2 yields all possible embedded surfaces’
(In the experiments the documents are the titles of papers drawn from ArXiv.)
The first step of the workflow classifies each statement according to basic academic field: Physics, Math, Biology and Computer Science. (Obviously there more fields than this, but this covered most of the collection we used to train the classifiers.) Once a sentence is classified as to topic it is then passed to the second stage of the workflow where it is classified as to subcategory. For example if a sentence is determined to belong to Biology, the subcategories that are recognized include Neuro, Cell Behavior, Genomics, Evolution, Subcellular, Health-Tissues&Organs and Molecular Networks. Physics sub areas are Astro, General Relativity and Quantum Gravity, Condensed Matter, High Energy, Mathematical Physics, Quantum mechanics and educational physics. Math is very hard, so the categories are simple: algebra, topology, analysis and other. The output from the DynamoDB table for this list of statements is shown in Figure 3 below.

Figure 3. Output from the sample document in an AWS DynamoDB table. The “cmain predict” column is the output of the first workflow phase and the “cpredicted” column is the output of the second phase of the workflow.
The AWS Lambda details.
A Lambda function is a very simple program that responds to an event, does some processing and exits. There is a complete command line interface to Lambda, but AWS has a very nice portal interface to build Lambda functions in a variety of standard languages. I found the web portal far superior to the command line because it gives you great debugging feedback and easy access to you function logs that are automatically generated each time your lambda function is invoked.
The example below is a very simple Python lambda function that waits for a document to arrive in a S3 storage bucket. When a document is placed in the bucket, the “lambda_handler” function is automatically invoked. In this simple example the function does three things. It grabs the name of the new S3 object and the bucket name. It then opens and reads the document (as text). If the document is not text, the system throws an error and the evidence is logged in the AWS CloudWatch log file for this function. In the last step, it saves the result in a DynamoDB table called “blanklambda”.
To make this work you need to assign the correct permissions policies to the lambda function. In this case we need access to S3, the DynamoDB and the basic Lambda execution role which includes permission to create the execution logs.
To tell the system which bucket to monitor you most go to the S3 bucket you want to monitor and add to the property called “Events”. Follow the instructions to a reference to your new Lambda function.

In our workflow example we used 5 lambda functions: a main topic classifier, and a classifier lambda function for each of the subtopics. It is trivial to make one Lambda function create an event that will trigger another. We send each document as a string json document encoding our list of statements.
The call function is
Lam = boto3.client("lambda",region_name="us-east-1")
resp = Lam.invoke(FunctionName="bio-function",
InvocationType="RequestResponse",
Payload=json.dumps(statementlist))
The “bio-function” Lambda code receives the payload as a text string and convert it back to a list.
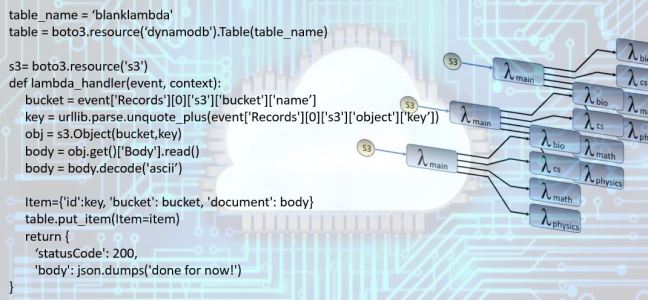
The workflow is pictured in Figure 4 below. When a document containing a list of statements lands in S3 it invokes an instance of the workflow. The main classifier lambda function invokes one instance each of the sub-classifiers and those four are all running concurrently. As illustrated in Figure 5, when a batch of 20 documents files land in s3 as many as 100 Lambda instances are running in parallel.

Figure 4. When a document list file lands in a specified S3 bucket it triggers the main lambda function which determines the general topic of each individual statement in the document list. The entire document is sent to each of the subtopic specialist lambda function that classifies the them into subtopics and places the result in the DynamoDB table.

Figure 5. As multiple documents land in S3 new instances of the workflow are run in parallel. A batch of 20 documents arriving all at once will generate 5*20 concurrent Lambda invocations.
AWS Lambda’s Greatest Limitation
Lambda functions are intended to be small and execute quickly. In fact, the default execution time limit is 3 seconds, but that is easily increased. What cannot be increased beyond a is the size of the package. The limit is 230Mbytes. To import python libraries that are not included in their basic package you must add “layers”. These layers are completely analogous to the layers in the Docker Unified File System. In the case of Python a layer is a zipped file containing the libraries you need. Unfortunately, there is only one standard library layer available for python 3 and that includes numpy and scipy. However, for our classification algorithms we need much more. A grossly simplified (and far less accurate) version of our classifier requires only sklearn, but that is still a large package.
It takes a bit of work to build a special Lambda layer. To create our own version of a Scikit learn library we turn to an excellent blog “How to create an AWS Lambda Python Layer” by Lucas. We were able to do this and have a layer that also included numpy. Unfortunately, we could not also include the model data, but we had the lambda instance dynamically load that data from S3. Because our model is simplified the load takes less than 1 second.
We tested this with 20 document files each containing 5 documents uniformly distributed over the 4 major topics. To understand the performance, we captured time stamps at the start and end of each instance of the main Lambda invocation. Another interesting feature of AWS Lambda execution is that the amount of CPU resource assigned to each instance of a function invocation is governed by the amount of memory you allocate to it. We tried our experiments with two different memory configurations: 380MB and 1GB.


Figure 6. Each horizontal line represents the execution of one Lambda workflow for one document. Lines are ordered by start time from bottom to top. The figure on the top show the results when the lambda memory configuration was set to 380MB. The figure on the bottom shows the results with 1GB of memory allocated (and hence more cpu resource).
The results are shown in Figure 6. There are two interesting points to note. First, the Lambda functions do not all start at the same time. The system seems to see that there are many events that were generated by S3 and after a brief start it launches an additional set of workflow instances. We don’t know the exact scheduling mechanism used. The other thing to notice is that the increase in memory (and CPU resource made a profound difference. While the total duration of each execution varied up to 50% between invocations the mean execution time for the large memory case was less than half that of the small memory case. In the 1GB case the system ran all 20 documents with a speed-up of 16 over the single execution case.
Having to resort to the simplified (and far less accurate) small classifier was a disappointment. An alternative, which in many ways is more appropriate for scientific workflows, is to use the Lambda functions as a the coordination and step-trigger mechanism and have the lambda functions invoke remote services to do the real computation. We ran instances of our full model as docker containers on the AWS LightSail platform and replaced the local invocations to the small classifier model with remote invocations to the full model as illustrated in Figure 7.

Figure 7. Invoking the remote classifiers as web services.
Results in this configuration were heavily dependent on the scale-out of each of the classifier service. Figure 8 illustrate the behavior.

Figure 8. Running 20 documents simultaneously pushed to S3 using remote webservice invocations for the classification step. Once again these are ordered from bottom to top by lambda start time.
As can be seen in the diagram the fastest time using a remote service for the full classifier was nearly three times faster than the small classifier running on the lambda function. However, the requests to the service caused a bottleneck which slowed down others. Our version of the service ran on two servers with a very primitive random scheduler for load balance. A better design would involve many more servers dynamically allocated to meet the demand.
Conclusion
Serverless computing is a technology that can simplify the design of some scientific workflow problems. If you can define the workflow completely in terms of the occurrence of specific triggers and the graph of the execution is a simple tree, it is easy to setup a collection of function that will be triggered by the appropriate events. If the events don’t happen, the workflow is not run and you do not have to pay for servers to host it. (The example from the previous section was designed, debugged on AWS over a few weeks and run many times. The total bill was about $5.)
There are two downsides to using serverless as part of a workflow execution engine.
- The complexity of handling a workflow consisting of a complex non-tree DAG is large. A possible solution to this was described in Figure 2 and the accompanying text. The task of compiling a general CWL specification into such a workflow manager/listener is non-trivial because CWL specification can be very complex.
- A topic that was only briefly discussed here is how to get a Lambda function to execute the actual application code at each step. In most workflow systems the work steps are command line invoked applications that take input files and produce output files. To invoke these from a serverless lambda function in the cloud requires that each worker application is rendered as a service that can be invoked from the lambda function. This can be done by containerizing the applications and deploying them in pods on Kubernetes or other microservice architecture as we illustrated in Figure 2. In other scenarios the worker applications may be submitted to a scheduler and run on a batch system.
While we only looked at AWS Lambda here we will next consider Azure functions. More on that later.
The source code for the lambda functions used here can be found at this GitHub repo
Note: there are various tricks to getting lambda to use larger ML libraries, but we didn’t go down that road. One good example is described in a blog by Sergei on Medium. Another look at doing ML on lambda is described by Alek Glikson in this 2018 article.

Reblogged this on Phill.
LikeLike